Normalization of Affymetrix DNA chip
D. Puthier
Inserm U1090/TAGC
Press F11 for full screen
Affymetrix DNA Chip
Technology |
 |
- In situ synthesis of oligonucleotides.
- Features.
- Cells: 24µm x 24µm
- ~ 1e7 oligos per cell
- ~ 4e5-1.5e6 probes per chip
Affymetrix DNA Chip
Probes |
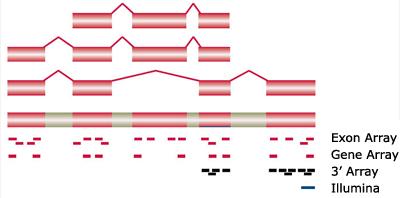 |
- Short oligonucleotides (25 mers)
- 3' arrays (3' IVT Expression Analysis)
- PM and MM
- 10-20 probes/gene
- Exon/Gene arrays (Whole-Transcript Expression Analysis)
- No more MM
- ~4 probes per exon, ~40 per gene (Exon Arrays)
- ~26 probes per gene (Gene arrays)
Affymetrix DNA Chip
Mismatch and perfect match |
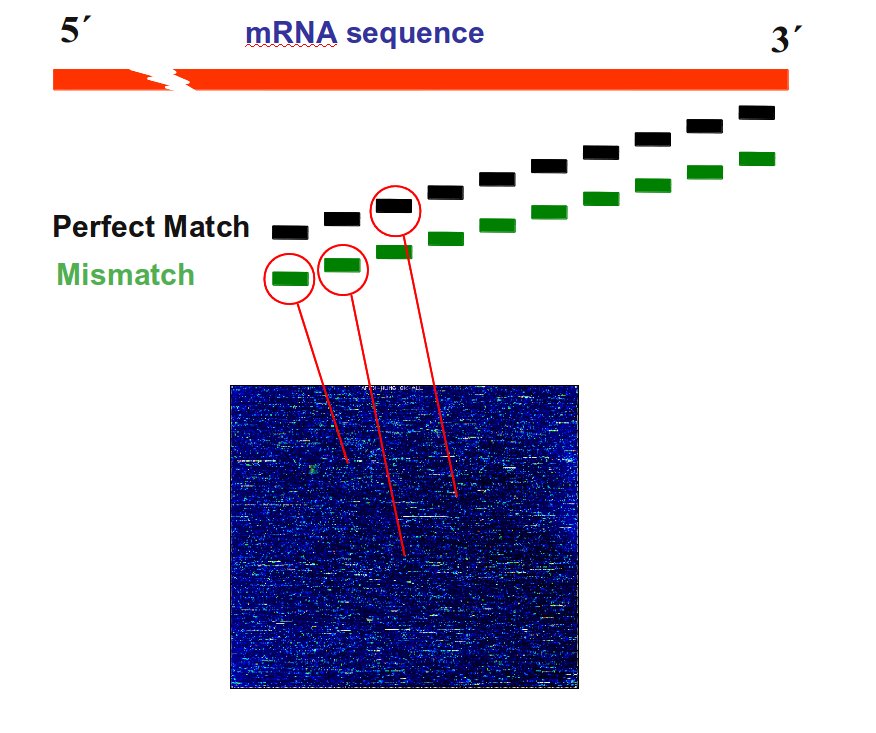 |
- Perfect Match (PM): specific probe
- Mismatch (MM): degenerated probe
- ProbePair: a pair of PM and MM
- ProbeSet: a set of x PM and MM targeting a gene (e.g; U12140_at)
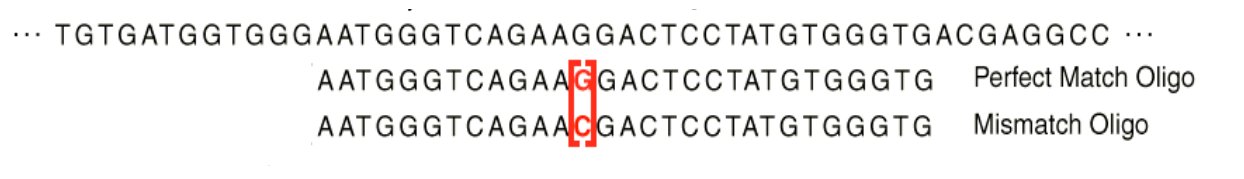

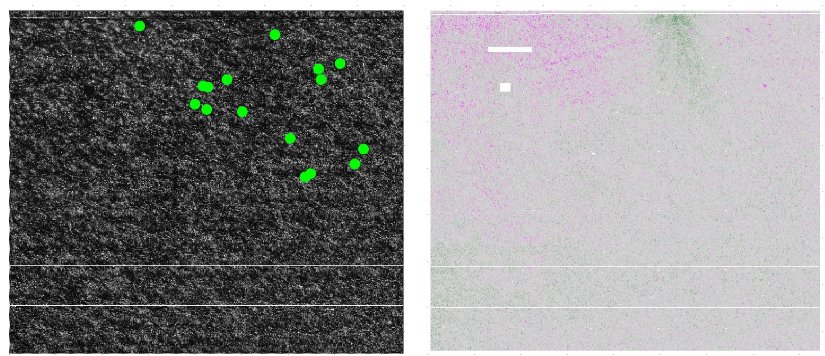
Normalization: main sources of artefact
- RNA quantity / quality
- Labelling efficiency
- Scanner parameters
- Local background

Normalizing columns of a numeric matrix
Standardization (set mean to 0 and standard deviation to 1 in each column)
- centering: substracting the mean from each value
- scaling: dividing the centered value by the standard deviation
- \(\ z \) is the standardized value (the z-score)
- \(\ m_{est} \) is a sample-based estimate of the population mean
- \(\ s_{est} \) is a sample-based estimate of the population standard deviation
Normalizing columns of a numeric matrix
Robust standardization (set median to 0 and median absolute deviation to 1 in each column)
- centering: substracting the median from each value
- scaling: dividing the centered value by the median absolute deviation
- \(\ z \) is the standardized value
- \(\ \tilde{m}_{est} \) is a sample-based estimate of the population median
- \(\ \tilde{s}_{est} \) is a sample-based estimate of the population mad
Normalizing columns of a numeric matrix
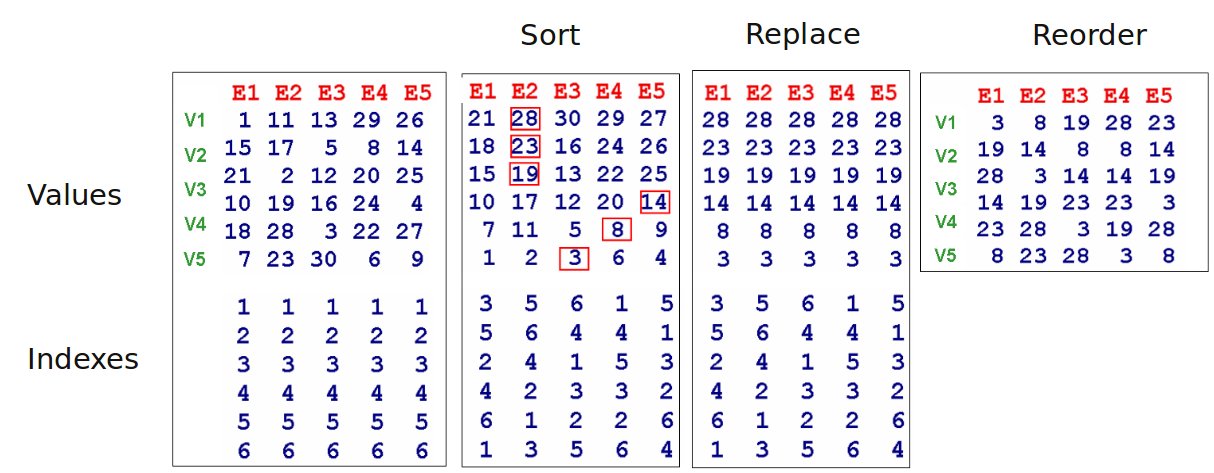
Quantile normalization
- Force columns to contain the same set of values
The RMA algorithm
Robust Multichip Average
The most popular algorithm for Affymetrix data normalization
- MM values are excluded from analysis
- Global background estimation and substraction
- Quantile normalization (probe level)
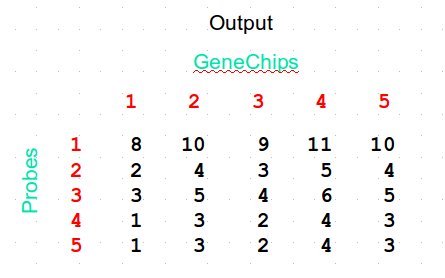
- Median Polish (summarization)
The RMA algorithm
Median polish algorithm
Motivations
- Probes may display higher or lower affinity for targets
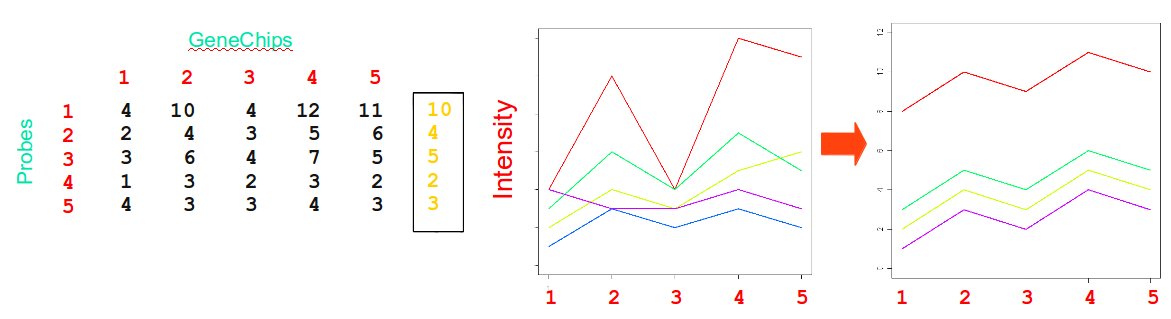
The RMA algorithm
Median polish algorithm
Procedure
- Compute a residual matrix by iteratively
- Substract the corresponding median from each row
- Substract the corresponding median from each column
- Stop if row medians and column medians are equal to zero
- Subtract the residual matrix from the initial matrix
- Compute the mean of each column to get summarized expression values